Project Details
Our OBJECTIVES
We aim to provide a synoptic assessment of seagrass-associated fish distributions along the Pacific coast, and determine a subset of fish species for which biogeography can be determined using eDNA metabarcoding. We aim to produce a highly-replicated assessment of seagrass community structure for answering questions about community assembly, assess climate sensitivity based on species distributional changes across time, and build a robust system of forecasting. We also aim to demonstrate feasibility and a community of practice for large-scale partnered multi-national environmental DNA observatory. We aim to follow FAIR guidelines for data sharing and to share protocols for others to join in and increase our monitoring potential.
|
Our ProcessAt each site, four eDNA water sample replicates are collected at 30cm depth, each separated by 10m, plus one negative field control. Bottles are capped, held on ice, and transported to a nearby on-shore processing station, where each sample is pumped through a sterile in-line positive-pressure filter and preserved in Longmire's buffer. Preserved samples are stored frozen and later processed to the Hakai Institute Quadra Centre for metabarcoding. Currently we are metabarcoding all samples at the 12S locus for identification of fish species. Contact us for detailed methods if you are interested.
|
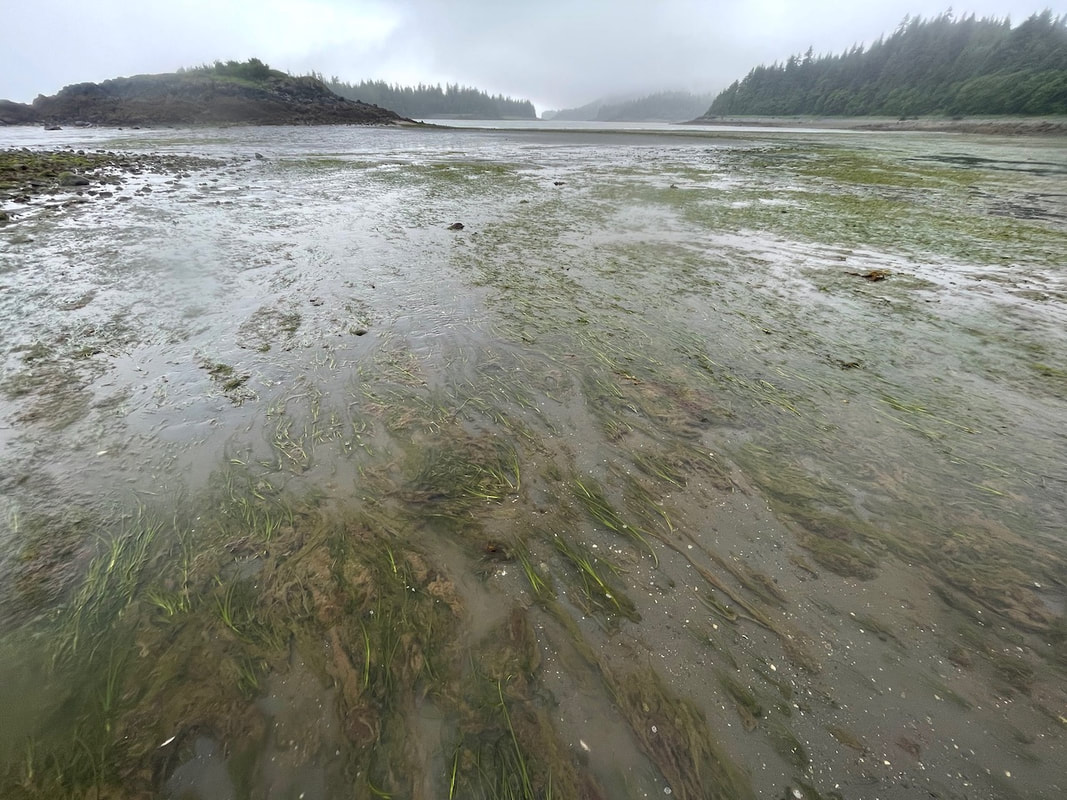
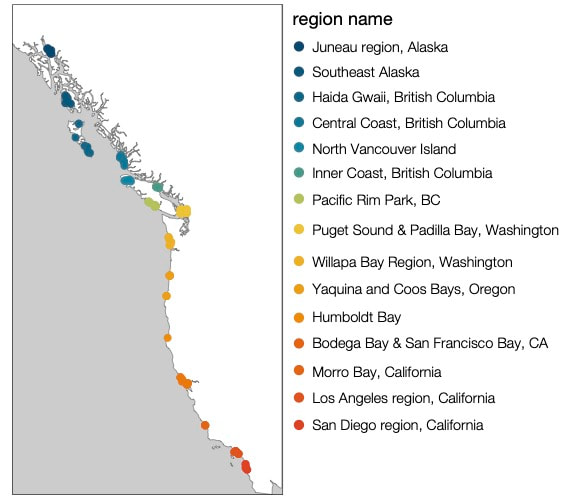
The SitesOur sites are nearshore seagrass habitats within regional 'nodes' that together span the West coast of North America. So far, we have focussed sampling within seagrass habitats, in order to standardize habitat type across space.
Our design is to sample 4 replicates within 5 sites within each region, with the goal of detecting the local regional fish fauna. Our regions are spatially balanced across four putative biogeographic breaks in order to answer questions about biogeographic turnover across space and time. |